Miscellaneous Utilities
Nicholas Mikolajewicz
2020-05-28
Source:vignettes/a05_Miscellaneous Utilities.Rmd
a05_Miscellaneous Utilities.RmdMiscellaneous Functions
There are several additional functions that are worthwhile exploring in the XcalRep package.
Data import and prep
Prior to analysis, import data and preprocess as described in the Getting Started Vignette.
# import EFP and QC1 imaging phantom data df.qc1 <-qc1 df.efp <- efp # omit water-mimetic resin sections df.qc1 <- dplyr::filter(df.qc1, section != 1) # remove water mimic section 1 df.efp <- dplyr::filter(df.efp, section != 4) # remove water mimic section 4 df.efp$section <- as.numeric(as.character(df.efp$section)) df.qc1$section <- as.numeric(as.character(df.qc1$section)) # combine datasets (ensure replicate sets aren't mixed) df.qc1$replicateSet <- paste("q", df.qc1$replicateSet, sep = "") df.efp$replicateSet <- paste("e", df.efp$replicateSet, sep = "") all.data <- bind_rows(df.qc1, df.efp) # create Calibration Object co <- createCalibrationObject(all.data) # get list of feature subsets to analyze analyze.these <- analyzeWhich(co, include.parameters = c("Tt.vBMD")) # preprocess/filter data co <- preprocessData(object = co, analyze.which = analyze.these, new.assay.name = "preprocessed.data", which.assay = "input")
1) Mean-Variance Relationship: meanVarPlot
The mean.var.plot function enables users to explore the relationship between measurement means and variance, and may inform which precision error is more appropriate to report. The precision error measure that remains relatively constant across all values is preferred, as it does not misrepresent reproducibility for extreme values.
This relationship can be examined as either a scatter plot
meanVarPlot(co, which.data = "uncalibrated", which.plot = "scatter", which.parameter = "Tt.vBMD")
#> Warning in meanVarPlot(co, which.data = "uncalibrated", which.plot =
#> "scatter", : precision errors calculated based on replicateSet grouping#> `geom_smooth()` using formula 'y ~ x'
#> `geom_smooth()` using formula 'y ~ x'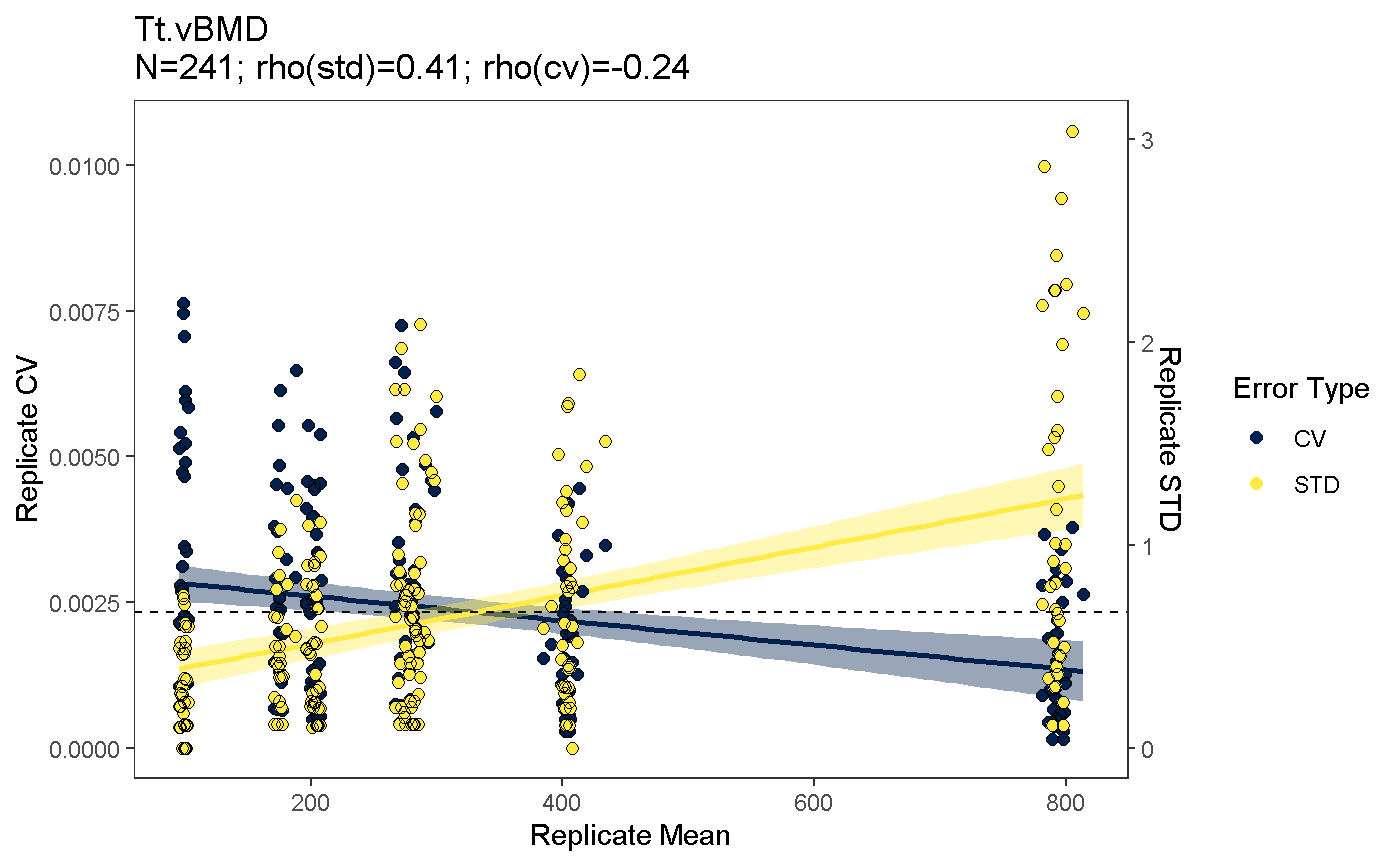
or boxplot
# requires troubleshooting # error message: Error in eval(predvars, data, env) : object 'section' not found # meanVarPlot(co, # which.data = "uncalibrated", # which.plot = "box", # which.parameter = "Tt.vBMD")
2) Utility Functions:
cloneAssay If users i) create an analysis “checkpoint” to which they can return at a later time or ii) diverge in analysis pipelines for the current dataset, cloneAssay will duplicate the specified (or current) Assay within the current Calibration Object.
# duplicate current assay co <- cloneAssay(co)
#>
#> 'preprocessed.data' was successfully clonedgetAssay(co, which.assays = "all")
#> [1] "input" "preprocessed.data" "preprocessed.data-copy"deleteAssay A specified assay can be deleted using the deleteAssay function.
# duplicate current assay co <- deleteAssay(co, which.assay = "preprocessed.data-copy")
#>
#> 'preprocessed.data-copy' was successfully deleted.